Population Genetics and Ancestry of the Haryanvi People
A deep dive
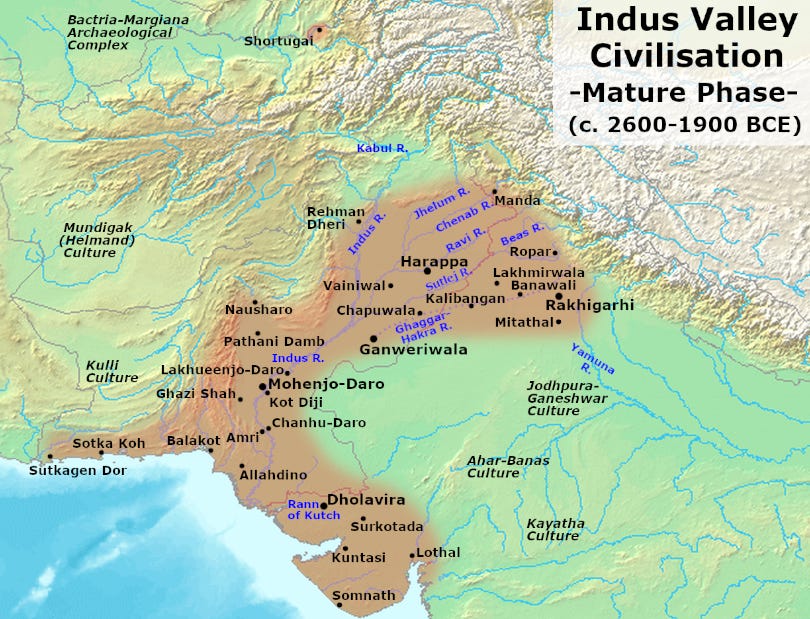
Haryana is a province/state in the northwestern parts of India, a region that was part of the Indus Valley Civilization.
Ancient DNA from a Sindhu-Saraswati (IVC) site at Rakhigarhi (Haryana) revealed a genetic profile derived mainly from an ancient population related to Iranian farmers and indigenous South Asian hunter-gatherers as per a study by Shinde et al.
It is generally surmised that the Indo-European (“Aryan”) genetic input arrived in Haryana after the peak of the Indus civilization. Subsequent migrations are supposed to have brought Steppe pastoralist ancestry into the region during the Bronze Age, which mixed with the local IVC gene pool. Present-day Haryanvi people thus carry a blend of these ancestries: a strong Ancestral North Indian (ANI) component (related to West Eurasian/steppe lineages) on top of the presumed ancient Indus Valley gene pool. This author’s modelling shows that the Rakhigarhi lady (ancient Harappan sample from Haryana, India) has significant amounts of Sarazm-related ancestry and Sarazm in turn has significant contribution from the Tutkaul hunter gatherer (TTK); this means Rakhigarhi has significant presence of the ANE (Ancient North Eurasian) ancestry component.
Genetic studies show that most Haryana populations fit along the typical North–South Indian genetic continuum, but certain groups exhibit greater affinity towards steppe ancestry. For example, the Ror community of Haryana stands out as genetically closer to groups northwest of India (e.g. ancient populations of the Indus-Swat region) than to many other Indians as per a paper on the genetics of modern Indus Valley populations from North India by Pathak, Kadian, Chaubey et al.
Researchers argue this is likely due to continuity of Bronze Age Indo-Aryan ancestry rather than recent foreign admixture. In other words, Haryana’s people have a deep lineage tracing back to early Vedic-era inhabitants, with relatively little later outside gene flow. Haryana’s gene pool is characterized by haplogroups common in North India. Paternal lineages frequently belong to Y-chromosome haplogroup R1a (associated with Indo-Aryan migrations) alongside others like J2, L, and Q. One extensive study of Haryanvi Jats found their male lineages split among at least nine haplogroups (R1a, J, L, Q, etc.), with ~90% falling into four major ones (J, L, Q, R) that they share with many other Indian communities. In contrast, another steppe-rich population from the state, the Rors have R1a, L1c, R2a, J2a and H1a as their most common male lineages.
This signals diverse ancient origins – not a single “founder” – for the population. Maternal mitochondrial DNA lineages in Haryana are equally mixed: roughly 40–50% of maternal haplogroups are West Eurasian-derived (types like U2, U7, W, R2) while the rest trace to deep Indian origins. Intriguingly, geneticists have argued for a male-biased influx of West Eurasian ancestry in the past but high steppe affinity groups from Haryana don’t show an elevated presence of R1a in their paternal gene pool.
For instance, the Ror and Jat populations show less than 40% of R1a and Q lineages in their respective paternal ancestries but autosomally a higher proportion of ancestral components are shared with ancient steppe herders. Overall, the people of Haryana today show a lot of genetic homogeneity with a clear signal of Indo-Aryan (steppe) heritage surviving since the times of the Indus Valley civilisation. In fact, Haryanvi groups like the Ror can serve as a proxy for the original “Ancestral North Indian” population component defined in population genetic models.
Agricultural and Dietary Adaptations
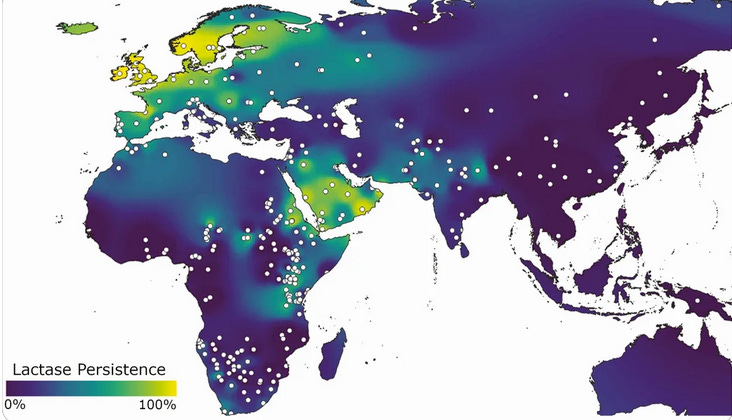
Lactose Tolerance – Dairy Adaptation: Haryana has a long tradition of dairy farming and milk consumption (the state is famed for its buffalo milk and ghee). Correspondingly, a significant fraction of its people have genetically adapted to digest lactose into adulthood. The ability to drink milk without discomfort, known as lactase persistence, is controlled by genetic variants. In most humans worldwide, the lactase enzyme turns off after weaning, causing lactose intolerance. But in populations with pastoral histories, mutations arose to keep lactase active. In India, lactase persistence is highest in the northwest. Notably, a study found the Ror community of Haryana has ~49% lactase persistence – meaning about half of Rors carry the gene variant that allows them to digest milk in adulthood. This is an exceptionally high rate for the Indian subcontinent (for comparison, some South Indian and tribal groups have as low as 0–5% lactase persistence) and South Central Asia. The map below illustrates this cline – North Indians (northwest) have far more lactose-tolerant individuals than the rest of India and South Central Asia.
The prevalent allele in Haryana is the same mutation (-13910*T in the LCT gene) found in Europeans, suggesting it likely spread with ancient herders. Thanks to this adaptation, dairy products form a staple of the Haryanvi diet – milk, yogurt (dahi), butter, and cheese – providing crucial nutrition. Those without the lactase-persistence allele can still consume milk, but often in fermented forms (curd, ghee) that are easier to digest. Overall, Haryana’s people show a clear genetic adaptation to an agro-pastoral diet, having one of the highest lactase persistence frequencies in India.
Comparisons with Other Populations
Within India: Genetically, Haryanvis are very much part of the North Indian mosaic. Their nearest relatives are the populations of neighboring states like Punjab, Rajasthan, Western Uttar Pradesh, and Himachal Pradesh. Studies consistently find minimal genetic distance between Haryana’s people and other Indo–Aryan speaking groups of the north. For example, an analysis of autosomal DNA (STR markers) showed that Brahmins from Haryana cluster closely with Brahmins from Rajasthan and even with Saraswat Brahmins of Himachal Pradesh. This indicates that caste and regional groupings in North India share a lot of ancestry across state lines – a Brahmin in Haryana may be genetically closer to a Brahmin in another state than to a non-Brahmin in his own state.
Haryanvis have one of the highest proportions of ANI, higher than Punjabis and most Pakistanis. They typically show stronger genetic affinity to populations of Pakistan and North India compared to southern castes or Austroasiatic-speaking tribes of India. It’s important to stress that these are average tendencies – within Haryana there is considerable genetic diversity, and individuals from different castes or locales might plot in slightly different positions in a genetic PCA space. The broad picture, however, places Haryana firmly in the North/Northwest Indian/Pakistani cluster. Interestingly, one genetic study used a statistical measure of shared genetic drift (Outgroup f3) and found that Haryanvi Rors share more drift with some European populations than do other South Asians.
This does not mean Haryanvis are Europeans; it reflects that both Haryanvis and Europeans inherit some common ancient ancestry (related to the Eurasian Steppe peoples and Early Neolithic farmers). In contrast, South Indian groups have less of that particular ancestry, so the genetic “distance” between a Haryanvi and, say, a Polish person is slightly smaller than that between a South Indian and a Polish person. In everyday terms, Haryanvis are genetically closer to Pakistanis and North Indians, moderately close to West Asians, North Africans and Central Asians, and more distant from East Asians or Africans.
Global Context and Unique Markers: When comparing Haryana’s genetics to global populations, a few features stand out. First, the Y-DNA haplogroup R1a is very common in Haryana (around 30–50% of male lineages in many communities) – this lineage is also prevalent in Eastern Europeans, Russians, and Central Asians. This shared haplogroup points to ancient links via Indo-European expansions. On the maternal side, haplogroups like M and U in Haryanvis show deep connections to other Indians/Pakistanis and West Eurasians respectively. However, no major lineage is absolutely unique to Haryana; it’s the combination and frequencies that characterize them. One could say the “genetic signature” of Haryana is: high frequency of lactase persistence allele (unique within India but similar to Europe), high frequency of blood group B (which is common in India but rarer in Europe/Americas), and presence of both India-specific lineages (like Y-haplogroup H or L) and Western Eurasian-related (R1a, J2). For instance, about 38% of Haryanvis have blood type B, whereas in Western countries type B is relatively uncommon. Conversely, the proportion of type O (universal donor) in Haryana (~31%) is lower than in many Western populations where O can exceed 40–50%.
Such differences in allele frequencies distinguish Haryanvis from other populations. Compared to East Asians, Haryanvis have almost none of the EDAR gene variants that cause shovel-shaped incisors and thicker hair – that variant is absent outside East Asia. Compared to Africans, Haryanvis have lighter skin on average due to nearly fixed “light skin” alleles similar to other West Eurasians.
Similarities: People from Haryana share a great deal of genetic heritage with other North Indians, Pakistanis, and to an extent Iranians, Arabs and Central Asians. They also share some ancient genetic lineages with Europeans. Physically and genetically, Haryanvis resemble other north-subcontinental groups – for example, a random person from Haryana might be virtually indistinguishable in DNA from someone in adjacent western Uttar Pradesh or eastern Punjab. This is why often genetic studies refer to a “North Indian cluster”. Culturally distinct neighboring populations (like Punjabis) are also genetically very close to Haryanvis due to continuous gene flow and similar ancestral mixtures. One large-scale Indian genomics project remarked that while North Indians and South Indians have distinct ancestral proportions, they are not completely isolated groups – many alleles are shared across India. In fact, that study noted no significant difference in certain drug-metabolizing gene frequencies between North (Haryana) and South Indians, underscoring substantial shared heritage.
References:
Frequency distribution of high risk alleles of CYP2C19, CYP2E1, CYP3A4 genes in Haryana population by Gulati, Yadav et al 2014
Distribution of ABO Blood Groups and RH (D) factor in Haryana by Puri et al 2016
The Genetic Ancestry of Modern Indus Valley Populations from Northwest India by Pathak, Chaubey, Kadian et al 2018
Genetic Diversity of Autosomal STR Markers in the Brahmin Population of Rajasthan and Haryana: Significance in Population and Forensic Genetics by Sharma, Sahajpal et al 2023
Y-STR Haplogroup Diversity in the Jat Population Reveals Several Different Ancient Origins by Mahal et al 2017
An Ancient Harappan Genome Lacks Ancestry from Steppe Pastoralists or Iranian Farmers by Shinde et al 2019
Haplotype Diversity of Mitochondrial DNA in the Jat Population of Haryana by Sharma et al 2023